This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Protein Interaction Networks
Because SLC6A4 is one protein in the vast network of proteins involved in neutrotransmitter uptake, signal transduction, and the nervous system itself, there are clearly many other proteins that work alongside it. No protein would be able to function correctly if it did not interact with other proteins somewhere in the rest of the body. For this reason, interaction network databases have been created to display the relationships between one protein to many others. When an interaction between two proteins is documented, it can lead to further understanding of the role those proteins play in their respective pathways [1].
Analysis
The STRING database was used to find the interaction network for the SLC6A4 protein. STRING uses the protein neighborhood, gene fusion, co-occurrence, co-expression, experiments, databases, textmining, and homolog information from published sources to map out relationships between different proteins, each represented by a different colored line. Interaction networks for the Homo sapiens, Rattus norvegicus, and Drosophila melanogaster were created and compared below.
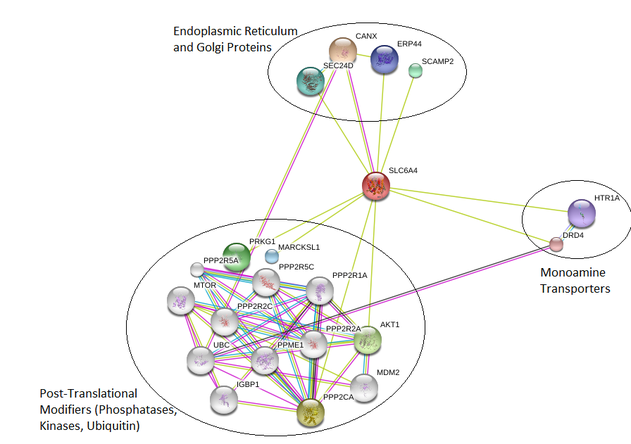
Figure 1: Interaction network for Homo sapiens SLC6A4 protein
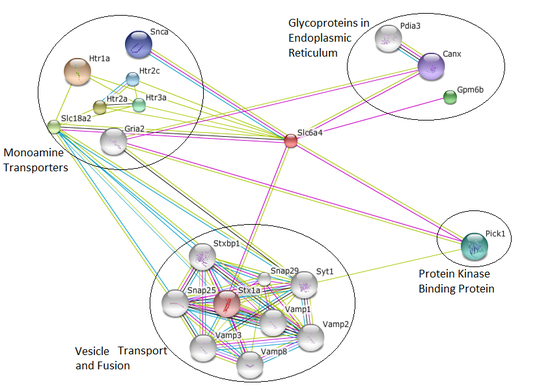
Figure 2: Interaction network for Rattus norvegicus Slc6a4 protein
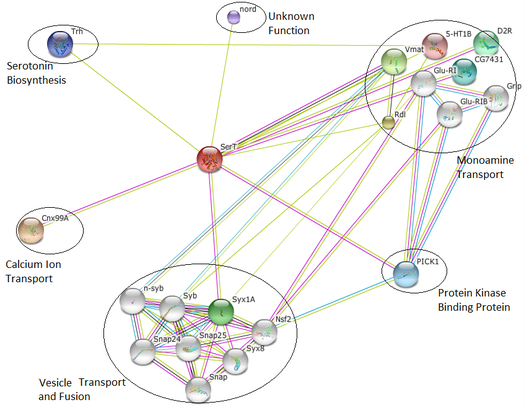
Figure 3: Interaction network for Drosophila melanogaster SerT protein
Discussion
Overall, the interaction networks for Homo sapiens, Rattus norvegicus, and Drosophila melanogaster showed information that would be expected given the known functions of serotonin transporters. All included a group with other monoamine transporters, such as dopamine. These are proteins that are all performing the same function, and many are within the same protein family because they are so closely related. There were also examples in each organism of calcium transporters, sometimes associated with the endoplasmic reticulum. Since serotonin release is triggered by an influx of sodium and chloride ions, it makes sense that transporters in a location with high stores of calcium such as the endoplasmic reticulum would be induced [2].
Human SLC6A4 has documented interaction with a number of protein kinases, phosphatases, and ubiquitin, all of which are involved in post-translational modification of proteins. In this case, suggested functions usually included localization of enzymes, substrate specificity, and catalytic activity. There are several post-translational modifications that have been identified in SLC6A4, so it is likely that these proteins are involved in not only this modification but also the modification of other proteins involved in serotonin transport. One protein kinase binding protein was documented in each of the rat and fly networks, PICK1, so there is probably post-translational modification in these organisms to some degree as well.
Rat and fly interaction networks show large groups of proteins related to vesicle transport. Serotonin is taken up into the presynaptic neuron and stored in vesicles until it needs to be released again, so there is a lot of reliance upon proper vesicle formation, transport, and docking [3]. These proteins ensure the serotonin is preserved properly after it interacts with the serotonin transporter.
There are a two protein relationships within Drosophila that are not found in either human or rat interaction networks. Based on the alternative structure of the fly SerT protein as opposed to the mammals, different interactions are more likely to be needed for the same functions to be carried out. One protein, Trh, is linked to biosynthesis of serotonin from tryptophan [4]. Serotonin clearly has an integral role in its serotonin transportation, as it is the substrate with which the serotonin transporter interacts. The second protein, nord, did not have a listed function so its relationship to the SerT serotonin transmitter could not be determined. It only has two GO terms associated with it, both in the biological processes category. They are "learning or memory" and "olfactory learning" [5]. More studies need to be carried out to find its place in the network of interactions, but the GO terms connected to the protein suggest that the relationship with serotonin transport may be linked to one of the effects of serotonin levels, such as memory and learning [6].
Human SLC6A4 has documented interaction with a number of protein kinases, phosphatases, and ubiquitin, all of which are involved in post-translational modification of proteins. In this case, suggested functions usually included localization of enzymes, substrate specificity, and catalytic activity. There are several post-translational modifications that have been identified in SLC6A4, so it is likely that these proteins are involved in not only this modification but also the modification of other proteins involved in serotonin transport. One protein kinase binding protein was documented in each of the rat and fly networks, PICK1, so there is probably post-translational modification in these organisms to some degree as well.
Rat and fly interaction networks show large groups of proteins related to vesicle transport. Serotonin is taken up into the presynaptic neuron and stored in vesicles until it needs to be released again, so there is a lot of reliance upon proper vesicle formation, transport, and docking [3]. These proteins ensure the serotonin is preserved properly after it interacts with the serotonin transporter.
There are a two protein relationships within Drosophila that are not found in either human or rat interaction networks. Based on the alternative structure of the fly SerT protein as opposed to the mammals, different interactions are more likely to be needed for the same functions to be carried out. One protein, Trh, is linked to biosynthesis of serotonin from tryptophan [4]. Serotonin clearly has an integral role in its serotonin transportation, as it is the substrate with which the serotonin transporter interacts. The second protein, nord, did not have a listed function so its relationship to the SerT serotonin transmitter could not be determined. It only has two GO terms associated with it, both in the biological processes category. They are "learning or memory" and "olfactory learning" [5]. More studies need to be carried out to find its place in the network of interactions, but the GO terms connected to the protein suggest that the relationship with serotonin transport may be linked to one of the effects of serotonin levels, such as memory and learning [6].
References
STRING: http://string-db.org/
[1] Alzate, O. (2010). Chapter 8: Protein Interaction Networks. In Neuroproteomics. Boca Raton, FL: Taylor & Francis. Retrieved March 22, 2015, from http://www.ncbi.nlm.nih.gov/books/NBK56024/
[2] Koch, G. (2005). The endoplasmic reticulum and calcium storage. Bioessays, 12(11), 527-531. Retrieved March 22, 2015, from http://onlinelibrary.wiley.com.ezproxy.library.wisc.edu/doi/10.1002/bies.950121105/abstract
[3] Neurotransmitter Release. (1998, January 1). Retrieved March 22, 2015, from http://web.williams.edu/imput/synapse/pages/II.html
[4] GH12537p. (n.d.). Retrieved March 22, 2015, from http://www.uniprot.org/uniprot/Q9W0K2
[5] AT02362p. (n.d.). Retrieved March 22, 2015, from http://www.uniprot.org/uniprot/Q8T4E5
[6] Symptoms of Depression. (n.d.). Retrieved March 21, 2015, from http://www.webmd.com/depression/guide/detecting-depression
Interaction Network Homo sapiens: http://string-db.org/newstring_cgi/show_network_section.pl?identifier=978773&additional_network_nodes=10&chemicalmode=-1&input_query_species=9606&interactive=yes&internal_call=1&limit=10&minprotchem=
0&network_flavor=evidence&previous_network_size=11&required_score=400&sessionId=QLpPQEnleLCB&targetmode=proteins&userId=2H0gObl_t3i_
Interaction Network Rattus noregicus: http://string-db.org/newstring_cgi/show_network_section.pl?identifier=1135739&additional_network_nodes=10&chemicalmode=-1&input_query_species=10116&interactive=yes&internal_call=1&limit=10&minprotchem=
0&network_flavor=evidence&previous_network_size=11&required_score=400&sessionId=QLpPQEnleLCB&targetmode=proteins&userId=2H0gObl_t3i_
Interaction Network Drosophila melanogaster: http://string-db.org/newstring_cgi/show_network_section.pl?identifier=623866&additional_network_nodes=10&chemicalmode=-1&input_query_species=7227&interactive=yes&internal_call=1&limit=10&minprotchem=
0&network_flavor=evidence&previous_network_size=11&required_score=400&sessionId=QLpPQEnleLCB&targetmode=proteins&userId=2H0gObl_t3i_
[1] Alzate, O. (2010). Chapter 8: Protein Interaction Networks. In Neuroproteomics. Boca Raton, FL: Taylor & Francis. Retrieved March 22, 2015, from http://www.ncbi.nlm.nih.gov/books/NBK56024/
[2] Koch, G. (2005). The endoplasmic reticulum and calcium storage. Bioessays, 12(11), 527-531. Retrieved March 22, 2015, from http://onlinelibrary.wiley.com.ezproxy.library.wisc.edu/doi/10.1002/bies.950121105/abstract
[3] Neurotransmitter Release. (1998, January 1). Retrieved March 22, 2015, from http://web.williams.edu/imput/synapse/pages/II.html
[4] GH12537p. (n.d.). Retrieved March 22, 2015, from http://www.uniprot.org/uniprot/Q9W0K2
[5] AT02362p. (n.d.). Retrieved March 22, 2015, from http://www.uniprot.org/uniprot/Q8T4E5
[6] Symptoms of Depression. (n.d.). Retrieved March 21, 2015, from http://www.webmd.com/depression/guide/detecting-depression
Interaction Network Homo sapiens: http://string-db.org/newstring_cgi/show_network_section.pl?identifier=978773&additional_network_nodes=10&chemicalmode=-1&input_query_species=9606&interactive=yes&internal_call=1&limit=10&minprotchem=
0&network_flavor=evidence&previous_network_size=11&required_score=400&sessionId=QLpPQEnleLCB&targetmode=proteins&userId=2H0gObl_t3i_
Interaction Network Rattus noregicus: http://string-db.org/newstring_cgi/show_network_section.pl?identifier=1135739&additional_network_nodes=10&chemicalmode=-1&input_query_species=10116&interactive=yes&internal_call=1&limit=10&minprotchem=
0&network_flavor=evidence&previous_network_size=11&required_score=400&sessionId=QLpPQEnleLCB&targetmode=proteins&userId=2H0gObl_t3i_
Interaction Network Drosophila melanogaster: http://string-db.org/newstring_cgi/show_network_section.pl?identifier=623866&additional_network_nodes=10&chemicalmode=-1&input_query_species=7227&interactive=yes&internal_call=1&limit=10&minprotchem=
0&network_flavor=evidence&previous_network_size=11&required_score=400&sessionId=QLpPQEnleLCB&targetmode=proteins&userId=2H0gObl_t3i_