This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Future Research
Obsessive compulsive disorder (OCD) is best described by the OCD Cycle. Obsessions start the cycle, consuming the thoughts of people with OCD and preventing them from focusing on anything else. These obsessions cause extreme anxiety, so to alleviate their stress and get rid of their obsessions, individuals with OCD turn to compulsive behavior. They might clean eight hours a day to deal with their fear of germs, or stand at their front door for an hour locking and relocking the house to make sure a robber will not be able to get in. While they are impractical and consume a huge part of their time, these compulsions temporarily give some relief from the obsessions and anxiety. Unfortunately, this relief is short-lived because within a very short time something else will usually trigger more obsessive thoughts and the cycle starts over again. This cycle is very hard to break and can lead to significant problems for people with OCD, including broken familial ties, isolation from society, and depression [1].
One gene that is known to be associated with OCD is the SLC6A4 serotonin transporter [2]. It is expressed in the membranes of presynaptic neurons and encodes a protein that causes reuptake of the neurotransmitter serotonin. The protein spans the neuronal membrane twelve times, as is characteristic of all monoamine transporters [3]. While there are several mutations in SLC6A4 that can cause its malfunction as a protein, one distinct missense mutation has been linked to OCD. At position 425, the isoleucine that is normally present gets swapped for a valine. One study found that this Ile425Val mutation caused SLC6A4 to be over-phosphorylated which in turn led to its over-expression, bringing in serotonin from the synapse in higher amounts than it is supposed to [4]. This phenomenon deserves greater exploration, so it is the basis of our specific aims. Rats were chosen as the model organism in these studies because they have been used as an OCD model for decades. They too exhibit anxiety-like behaviors, such as ritualistic grooming, and are highly conserved SLC6A4 homologs [5]. Their similarities to humans will make our results more reliable in the long-term than other model organisms.
One gene that is known to be associated with OCD is the SLC6A4 serotonin transporter [2]. It is expressed in the membranes of presynaptic neurons and encodes a protein that causes reuptake of the neurotransmitter serotonin. The protein spans the neuronal membrane twelve times, as is characteristic of all monoamine transporters [3]. While there are several mutations in SLC6A4 that can cause its malfunction as a protein, one distinct missense mutation has been linked to OCD. At position 425, the isoleucine that is normally present gets swapped for a valine. One study found that this Ile425Val mutation caused SLC6A4 to be over-phosphorylated which in turn led to its over-expression, bringing in serotonin from the synapse in higher amounts than it is supposed to [4]. This phenomenon deserves greater exploration, so it is the basis of our specific aims. Rats were chosen as the model organism in these studies because they have been used as an OCD model for decades. They too exhibit anxiety-like behaviors, such as ritualistic grooming, and are highly conserved SLC6A4 homologs [5]. Their similarities to humans will make our results more reliable in the long-term than other model organisms.
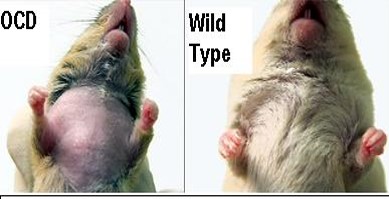
Figure 1: Behavioral example of OCD-like symptoms in rats. This is often characterized by a ritualistic grooming behavior in which they end up uncontrollably tearing out the fur on their chests. This is not seen in wild-type rats.
Though research is ongoing, there is still a long way to go and much more to discover about OCD. This is a polygenic disorder, with much more than SLC6A4 contributing to the symptoms. There are also many environmental factors that can affect the severity of symptoms or how they manifest. For this reason, we have chosen to look further into the complex protein interactions that lead to OCD with the hope that our findings will not only improve the lives of individuals with OCD but also those with related disorders across the spectrum of anxiety and depressive disorders. Based on the findings concerning phosphorylation of the SLC6A4 Ile425Val mutant, we hypothesize that the Ile425Val mutantation increases the expression of certain protein kinases, and those in turn cause alternative phosphorylation in other proteins in the body. These proteins would likely impact the development of OCD in conjunction with SLC6A4 and bring more understanding of the polygenic nature of the disorder.
Aim 1
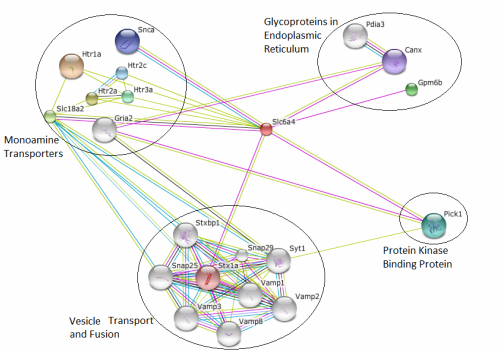
This aim is designed to identify novel and conserved SLC6A4 interacting proteins. While there is a fairly extensive interaction map in STRING for the rattus norvegicus SLC6A4 protein, only a single interaction is found with a protein kinase as shown in the STRING map below. The use of tandem affinity purification (TAP) tags will expose SLC6A4-protein interactions that have not have been seen before because of the extreme sensitivity of the technique. These interactions will be compared to those found in STRING, and any novel interacting proteins will be entered into BLAST to find human homologs. If there are human homologs, it is possible that these also contribute to the symptoms seen when SLC6A4 is mutated. Any new interacting proteins are potential candidates for further study.
It would be expected that several more protein kinases will be found to interact with SLC6A4 than are currently documented in STRING. The predicted phosphorylation sites for the rattus norvegicus SLC6A4 protein are extensive, suggesting phosphorylation by much more than a single kinase.
Aim 2
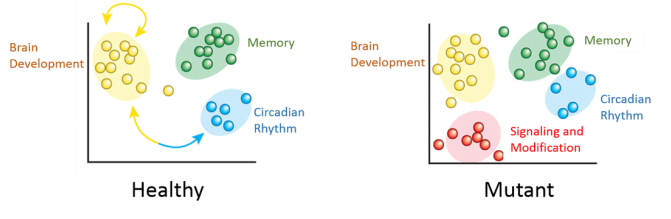
With more extensive interactions discovered in aim 1, we now want to identify genes that are misregulated in SLC6A4 mutant rats containing the Ile425Val mutation. These mutants will be compared to wild-type rats. Neuronal tissue will be collected from both groups and used to assess gene expression levels using RNA-seq. Based on the results of the previous aim, it would be expected that several of the newly discovered interacting protein kinases will be upregulated in the mutant rats.
This will be confirmed by entering differentially expressed genes into the Gene Ontology Consortium (GO) and grouping them based on their biological processes. Protein kinases generally function in signaling and protein modification, so it is likely that several genes that are upregulated will be grouped into these categories. Often, protein kinases do not phosphorylate a single protein, so those that interact with SLC6A4 probably interact with other proteins as well. This rationale will be tested in our final aim.
Aim 3
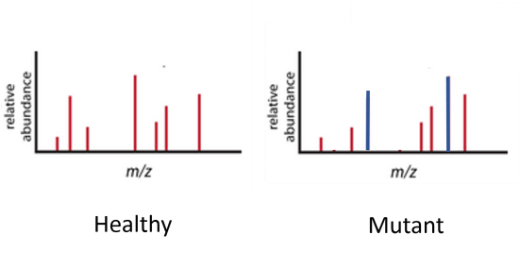
With the expectation that one protein kinase will phosphorylate more than one protein, we will identify alternatively phosphorylated proteins due to interactions with the kinases from the previous aim. Again neuronal tissue samples from healthy and SLC6A4 mutant rats will be collected. The samples will be tested with tandem mass spectrometry, allowing all proteins to be identified. The spectra for each sample will differ whenever an extra phosphate has been added to the protein, moving the peak significantly to the right.
It would be expected that those proteins with an increase in phosphorylation sites in the mutant sample are associating with the kinases identified in the previous aim. These proteins are good candidates for future studies on OCD because they are likely contributing to some of the characteristic behavioral changes experienced with the disorder.
Future Directions
The hopeful outcome of these experiments is to determine kinases that are affected by a missense mutation in the SLC6A4 protein that has been associated with OCD. Those kinases will likely affect other proteins, which could contribute to the diverse set of symptoms that characterize OCD. Each of those proteins will be studied in the future to determine how they interact with each other. Very similar studies to those detailed above can be performed for each of these proteins so that ultimately a complex understanding of their interactions as a network can be clarified. Only knowledge of the network as a whole will provide the information necessary to develop treatments that encompass all the symptoms of OCD. After the system is understood, gene therapies and drugs could be developed to target the specific genes that cause the most problems and give some relief to the people affected by this debilitating disorder.
In addition, it has been found that OCD and other anxiety disorders are genetically similar to depressive disorders [6]. When the networks of interaction are understood for OCD, the knowledge gained can likely be applied to the study of other disorders. Not only will treatments be made available for individuals with OCD, but for millions of others with related issues such as bipolar disorder or general depressive disorder.
In addition, it has been found that OCD and other anxiety disorders are genetically similar to depressive disorders [6]. When the networks of interaction are understood for OCD, the knowledge gained can likely be applied to the study of other disorders. Not only will treatments be made available for individuals with OCD, but for millions of others with related issues such as bipolar disorder or general depressive disorder.
| steinbrinkfinaltalk5-17-15.pdf | |
| File Size: | 1612 kb |
| File Type: | |
| steinbrinkupdatedfinalpresentation.pptx | |
| File Size: | 5894 kb |
| File Type: | pptx |
| steinbrinkfinalpresentation.pptx | |
| File Size: | 5500 kb |
| File Type: | pptx |
| steinbrinkdraft2presentation.pptx | |
| File Size: | 6096 kb |
| File Type: | pptx |
| steinbrinkdraft1presentation.pptx | |
| File Size: | 6369 kb |
| File Type: | pptx |
References
[1] Kring, A., & Johnson, S. (2013).
Obsessive-Compulsive and Related Disorders. In Abnormal Psychology
(12th ed. ed., DSM-5 Update, pp. 202-213). Hoboken, N.J.: John Wiley & Sons.
[2] Nestadt, G., Grados, M., & Samuels, J. (2010). Genetics of OCD. Psychiatr Clin North Am, 33(1), 141-158. Retrieved April 15, 2015, from http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2824902/
[3] SLC6A4 - Sodium-dependent serotonin transporter - Homo sapiens (Human). (n.d.). Retrieved February 19, 2015, from http://www.uniprot.org/uniprot/P31645
[4] Ramamoorthy, S., Shippenberg, T., & Jayanthi, L. (2011). Regulation of monoamine transporters: Role of transporter phosphorylation. Pharmacology & Therapeutics, 129(2), 220-238. Retrieved April 14, 2015, from http://www.sciencedirect.com/science/article/pii/S0163725810001932
[5] Kreiss, D., Coffman, C., Fiacco, N., Granger, J., Helton, B., Jackson, J., . . . Zimmer, B. (2013). Ritualistic Chewing Behavior induced by mCPP in the rat is an animal model of Obsessive Compulsive Disorder. Pharmacology Biochemistry and Behavior, 104, 119-124. Retrieved February 22, 2015, from http://www.sciencedirect.com.ezproxy.library.wisc.edu/science/article/pii/S0091305713000105
[6] Pallanti, S., Grassi, G., Sarrecchia, E., Cantisani, A., & Pellegrini, M. (2011). Obsessive–Compulsive Disorder Comorbidity: Clinical Assessment and Therapeutic Implications. Front. Psychiatry Frontiers in Psychiatry, 2(70), 1-11. Retrieved May 17, 2015, from http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3243905/
STRING: http://string-db.org/
BLAST: http://blast.ncbi.nlm.nih.gov/Blast.cgi
Gene Ontology Consortium: http://geneontology.org/
Pictures top to bottom:
Rat OCD Symptoms: http://newswise.com/articles/nobel-winner-ties-mental-illness-to-immune-defect
Rat STRING network: http://string-db.org/newstring_cgi/show_network_section.pl?identifier=1135739&additional_network_nodes=10&chemicalmode=-1&input_query_species =10116&interactive=yes&internal_call=1&limit=10&minprotchem=0&network_flavor=evidence&previous_network_size=11&required_score=400&sessionId=QLpPQEnleLCB&targetmode=proteins&userId=2H0gObl_t3i__
RNA-Seq: http://www.nature.com.ezproxy.library.wisc.edu/nmeth/journal/v11/n1/full/nmeth.2764.html
Tandem Mass Spec: http://web.science.uu.nl/developmentalbiology/boxem/interaction_mapping.html
[2] Nestadt, G., Grados, M., & Samuels, J. (2010). Genetics of OCD. Psychiatr Clin North Am, 33(1), 141-158. Retrieved April 15, 2015, from http://www.ncbi.nlm.nih.gov/pmc/articles/PMC2824902/
[3] SLC6A4 - Sodium-dependent serotonin transporter - Homo sapiens (Human). (n.d.). Retrieved February 19, 2015, from http://www.uniprot.org/uniprot/P31645
[4] Ramamoorthy, S., Shippenberg, T., & Jayanthi, L. (2011). Regulation of monoamine transporters: Role of transporter phosphorylation. Pharmacology & Therapeutics, 129(2), 220-238. Retrieved April 14, 2015, from http://www.sciencedirect.com/science/article/pii/S0163725810001932
[5] Kreiss, D., Coffman, C., Fiacco, N., Granger, J., Helton, B., Jackson, J., . . . Zimmer, B. (2013). Ritualistic Chewing Behavior induced by mCPP in the rat is an animal model of Obsessive Compulsive Disorder. Pharmacology Biochemistry and Behavior, 104, 119-124. Retrieved February 22, 2015, from http://www.sciencedirect.com.ezproxy.library.wisc.edu/science/article/pii/S0091305713000105
[6] Pallanti, S., Grassi, G., Sarrecchia, E., Cantisani, A., & Pellegrini, M. (2011). Obsessive–Compulsive Disorder Comorbidity: Clinical Assessment and Therapeutic Implications. Front. Psychiatry Frontiers in Psychiatry, 2(70), 1-11. Retrieved May 17, 2015, from http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3243905/
STRING: http://string-db.org/
BLAST: http://blast.ncbi.nlm.nih.gov/Blast.cgi
Gene Ontology Consortium: http://geneontology.org/
Pictures top to bottom:
Rat OCD Symptoms: http://newswise.com/articles/nobel-winner-ties-mental-illness-to-immune-defect
Rat STRING network: http://string-db.org/newstring_cgi/show_network_section.pl?identifier=1135739&additional_network_nodes=10&chemicalmode=-1&input_query_species =10116&interactive=yes&internal_call=1&limit=10&minprotchem=0&network_flavor=evidence&previous_network_size=11&required_score=400&sessionId=QLpPQEnleLCB&targetmode=proteins&userId=2H0gObl_t3i__
RNA-Seq: http://www.nature.com.ezproxy.library.wisc.edu/nmeth/journal/v11/n1/full/nmeth.2764.html
Tandem Mass Spec: http://web.science.uu.nl/developmentalbiology/boxem/interaction_mapping.html