This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Post-Translational Modifications
Even after mRNA has been translated into protein, it is not completely ready to perform its function in the cell. Instead there are a number of post-translational modifications that can occur that alter how the protein interacts with other proteins, signal where it is supposed to go in the cell, or even mark it for degradation. The most common of these modifications is phosphorylation, though it is only one of the dozens that could potentially be added. Phosphorylation sites are present on many proteins and play an important role in both activating and inactivating proteins, as well as facilitating protein-protein or protein-DNA interactions [1].
Analysis
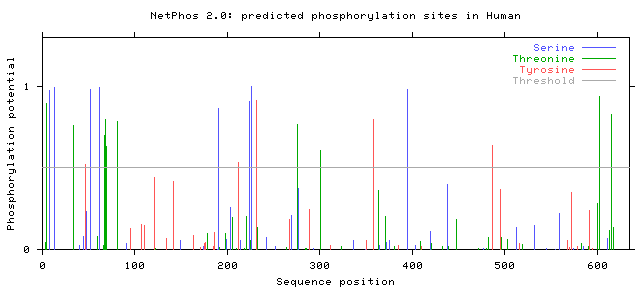
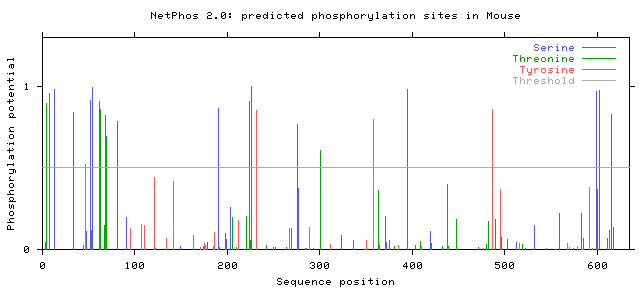
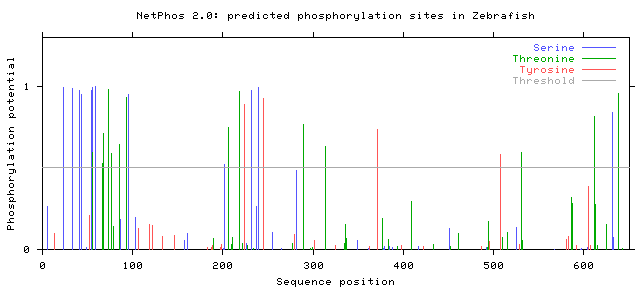
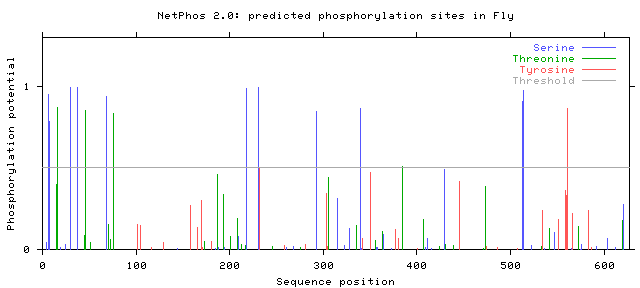
The NetPhos 2.0 database was used to find potential phosphorylation sites on serine, threonine, and tyrosine residues in humans, mice, zebrafish, and flies. The protein sequence from BLAST was entered, and the graphs of the potential phosphorylation sites that were given as the output are shown below.
Note that only those sites where the phosphorylation potential exceeds the threshold are likely to be phosphorylated.
Discussion
As the data above suggests, phosphorylation plays a significant role in the post-translational modification of SLC6A4 and likely to the ultimate function of the protein. This corroborates other studies that have been done on the phosphorylation of serotonin transporters, including the procedure outlined by Ramammorthy, Shippenberg, and Jayanthi. Among others, they used an SLC6A4 mutant protein with an amino acid sequence change at position 425 from an isoleucine to a valine (Ile425Val). This change led to constitutive phosphorylation at position 425 such that the protein was constantly activated and there was an increase in serotonin uptake. While NetPhos 2.0 only predicts phosphorylation sites at serine, threonine, and tyrosine, it is very possible that other mutations similar to the Ile425Val exchange occur at some of these predicted sites as well.
Phosphorylation potentials appear to be very similar between the organisms tested. This is not a surprise when protein homology is taken into account. Conservation is very high between organisms with SLC6A4 homologs, even those distantly related such as humans and flies, and therefore it makes sense that phosphorylation sites remain similar as well. All organisms show many more predicted serine and threonine sites than tyrosine, with distinct clusters forming between amino acids 1-100, 200-250, and from 600 to the end of the protein. The first and last of these segments are parts of the protein that are in the cytoplasm, while the segment from 200-250 is outside the cell in the synapse [3]. In either case, these are all parts of the protein that are directly interacting with the environment and not spanning the neural membrane. Since phosphorylation sites are generally areas where the protein is interacting with other proteins or molecules, either for signaling purposes or protein regulation, exposure to the cytoplasm or synapse is critical because this is where those interactions will occur [1].
Phosphorylation potentials appear to be very similar between the organisms tested. This is not a surprise when protein homology is taken into account. Conservation is very high between organisms with SLC6A4 homologs, even those distantly related such as humans and flies, and therefore it makes sense that phosphorylation sites remain similar as well. All organisms show many more predicted serine and threonine sites than tyrosine, with distinct clusters forming between amino acids 1-100, 200-250, and from 600 to the end of the protein. The first and last of these segments are parts of the protein that are in the cytoplasm, while the segment from 200-250 is outside the cell in the synapse [3]. In either case, these are all parts of the protein that are directly interacting with the environment and not spanning the neural membrane. Since phosphorylation sites are generally areas where the protein is interacting with other proteins or molecules, either for signaling purposes or protein regulation, exposure to the cytoplasm or synapse is critical because this is where those interactions will occur [1].
References
NetPhos 2.0: http://www.cbs.dtu.dk/services/NetPhos/
[1] Phosphorylation. (n.d.). Retrieved April 12, 2015, from https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/phosphorylation.html
[2] Ramamoorthy, S., Shippenberg, T., & Jayanthi, L. (2011). Regulation of monamine transporters: Role of transporter phosphorylation. Pharmacology & Therapeutics, 129(2), 220-238. Retrieved April 14, 2015, from http://www.sciencedirect.com/science/article/pii/50163725810001932
[3] InterPro. (n.d.). Retrieved April 28, 2015, from http://www.ebi.ac.uk/interpro/sequencesearch/iprscan5-S20150428-060500-0856-26021861-oy
[1] Phosphorylation. (n.d.). Retrieved April 12, 2015, from https://www.lifetechnologies.com/us/en/home/life-science/protein-biology/protein-biology-learning-center/protein-biology-resource-library/pierce-protein-methods/phosphorylation.html
[2] Ramamoorthy, S., Shippenberg, T., & Jayanthi, L. (2011). Regulation of monamine transporters: Role of transporter phosphorylation. Pharmacology & Therapeutics, 129(2), 220-238. Retrieved April 14, 2015, from http://www.sciencedirect.com/science/article/pii/50163725810001932
[3] InterPro. (n.d.). Retrieved April 28, 2015, from http://www.ebi.ac.uk/interpro/sequencesearch/iprscan5-S20150428-060500-0856-26021861-oy