This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Domains
Domains are portions of proteins that that are able to perform a specific function or form a distinct structure. A single domain may be found in many types of proteins, and there may be more than one of the same domain within a single protein. The identification of domains is one step in the process of grouping proteins into families that all perform a similar function. If one domain is known within a protein, part of its function can often be determined which is useful for newly discovered or uncharacterized proteins [1].
Analysis
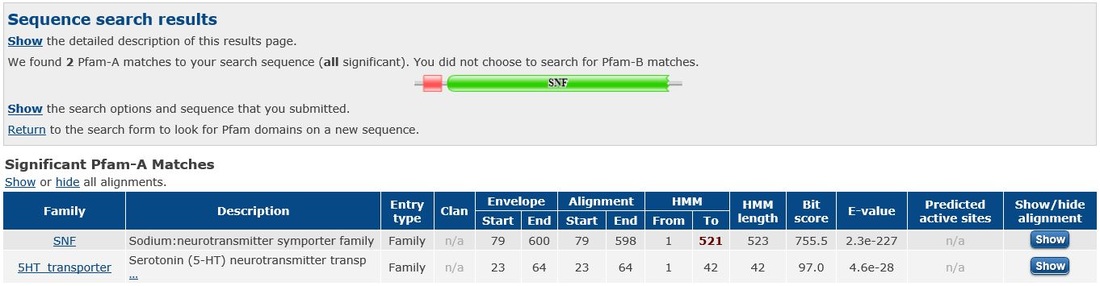
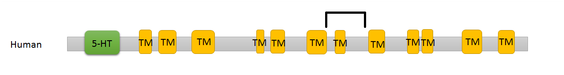
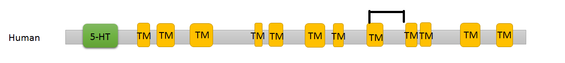
Domains within the SLC6A4 protein were found using Pfam and InterPro. Pfam displayed two domains for humans, both the 5-HTT domain and a sodium neurotransmitter symporter family domain. The 5-HTT domain is the domain in which the 5-HTT L and S alleles are based. The sodium transmitter symporter family domain is a domain found in all SLC6 proteins and includes the twelve transmembrane segments that are characteristic of this family of proteins.
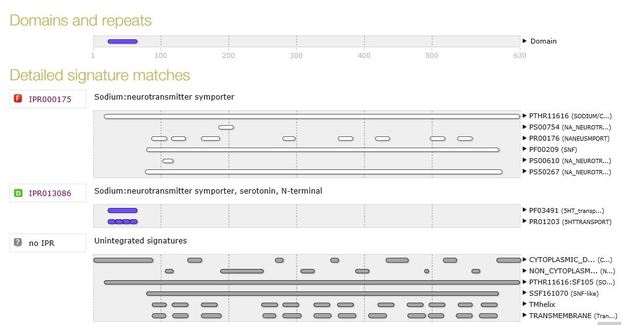
A search within InterPro provided similar results for the human domains, but each of the transmembrane segments were given rather than a single broad symporter family domain.
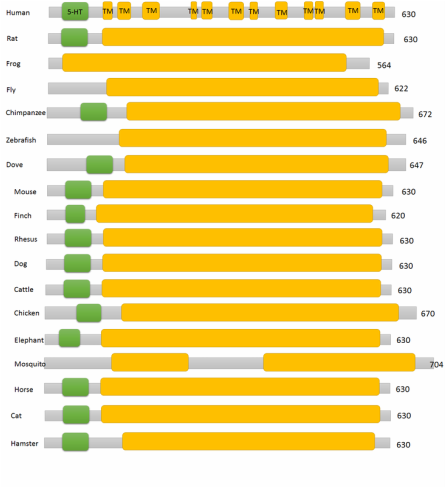
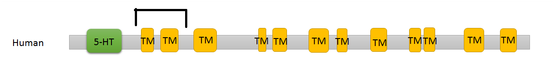
Results of the seventeen SLC6A4 homologs are listed below. The twelve human transmembrane segments are depicted individually, but all other homologs are shown as the sodium transmitter symporter family domain for simplicity.
Motifs
There are sometimes sequences within proteins that lend them a specific function. These sequences are called motifs, and they can be used to determine certain functions of unknown or uncharacterized proteins. They can also be used to group many proteins together into families that have similar functions. Sometimes motifs are within domains, but this is not always true and should not be assumed when doing an analysis of motifs [2].
Analysis
MEME was used to find three motifs common to the seventeen homologs as well as the Homo sapiens SLC6A4 protein. Protein sequences were entered using a FASTA format and saved as a plain text file. A sample document is provided below.
Motif 1
The first motif is located between the 98 and 147 amino acids in Homo sapiens. This corresponds to the amino acid sequence covering the first two transmembrane segments.
Motif 2
Motif 2 is located between amino acids 351 and 400 in Homo sapiens. This also corresponds to an area of the sodium transmitter symporter family domain, specifically the seventh transmembrane segment.
Motif 3
The final motif analyzed is located between the 414 and 463 amino acids in Homo sapiens. This spans the eighth transmembrane segment.
Motifs and Domains Discussion
Domains within the SLC6A4 protein of various homologs are highly conserved in most cases. The most notable difference is the lack of 5-HTT domains within the organisms that are presumably least related to Homo sapiens (insects, fish, and amphibians). This suggests that the domain was likely added at some time in evolution between the emergence of amphibians and mammals. Drosophila melanogaster SerT is known to be less affected by certain antidepressants and is less dependent on the influx of chloride ions than the human SLC6A4. Some of these characteristics may be rooted in the missing 5-HTT domain [3]. It also indicates that the L and S alleles found in the 5-HTT domain in humans is a non-issue in these organisms and they do not experience the symptoms of OCD through the same mechanism that mammals do. No studies were found addressing this topic.
Common motifs between all homologs tested were all found within the sodium transmitter symporter family domain. Since serotonin molecules have to be transported in all of these organisms, it makes sense that there would be consensus sequences to allow its passage with efficiency regardless of the organism. The serotonin molecule doesn't change, so its transporter is not able to mutate much without it affecting the ability of serotonin to pass through the membrane. Consensus sequences were not found within the 5-HTT region, almost certainly because four of the homologs did not express this domain in their respective serotonin transporter proteins. This part of the protein is not highly conserved enough to contain a motif within it that applies to the wide variety of organisms included in the analysis.
Common motifs between all homologs tested were all found within the sodium transmitter symporter family domain. Since serotonin molecules have to be transported in all of these organisms, it makes sense that there would be consensus sequences to allow its passage with efficiency regardless of the organism. The serotonin molecule doesn't change, so its transporter is not able to mutate much without it affecting the ability of serotonin to pass through the membrane. Consensus sequences were not found within the 5-HTT region, almost certainly because four of the homologs did not express this domain in their respective serotonin transporter proteins. This part of the protein is not highly conserved enough to contain a motif within it that applies to the wide variety of organisms included in the analysis.
References
Pfam: http://pfam.xfam.org/
InterPro: http://www.ebi.ac.uk/interpro/
MEME: http://meme.nbcr.net/meme/tools/meme
[1] Train online. (n.d.). Retrieved March 22, 2015, from http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains
[2] D'haeseleer, P. (2006). What are DNA sequence motifs? Nature Biotechnology, 24, 423-425. Retrieved March 22, 2015, from http://www.nature.com/nbt/journal/v24/n4/full/nbt0406-423.html
[3] Martin, C., & Krantz, D. (2014). Drosophila melanogaster as a genetic model system to study neurotransmitter transporters. Neurochemistry International, 73, 71-88. Retrieved March 22, 2015, from http://www.sciencedirect.com.ezproxy.library.wisc.edu/science/article/pii/S0197018614000709
InterPro: http://www.ebi.ac.uk/interpro/
MEME: http://meme.nbcr.net/meme/tools/meme
[1] Train online. (n.d.). Retrieved March 22, 2015, from http://www.ebi.ac.uk/training/online/course/introduction-protein-classification-ebi/protein-classification/what-are-protein-domains
[2] D'haeseleer, P. (2006). What are DNA sequence motifs? Nature Biotechnology, 24, 423-425. Retrieved March 22, 2015, from http://www.nature.com/nbt/journal/v24/n4/full/nbt0406-423.html
[3] Martin, C., & Krantz, D. (2014). Drosophila melanogaster as a genetic model system to study neurotransmitter transporters. Neurochemistry International, 73, 71-88. Retrieved March 22, 2015, from http://www.sciencedirect.com.ezproxy.library.wisc.edu/science/article/pii/S0197018614000709