This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Transcriptomics
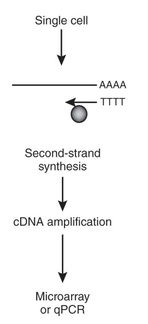
Transcriptomics involves the study of mRNA transcripts in target cells, between cell or tissue types, or between wild-type and mutant cells to determine differential expression of genes. The mRNA is collected and reverse transcriptase used to create a cDNA library. These libraries are useful as opposed to normal DNA libraries because they do not contain any intron segments. Instead they are DNA sequences that directly transcribe mRNA transcripts without modifications [1].
cDNA libraries are the key to transcriptomic microarrays. The cDNA is immobilized on a solid surface and a solution of labeled mRNA molecules from the target cells is added. mRNA from the control will be labeled one color and the test sample will be labeled another. As a result, locations on the solid surface that fluoresce the color of the test mRNA label are said to show increased expression of the corresponding gene compared to the control. Alternatively, if the spot fluoresces the color of the control there is under-expression of the gene in the test sample. When expression is seen in both subjects, a third color is produced [1]. Once the colors are analyzed and linked to their respective genes, grouping techniques such as GO can be used to determine what GO terms are present in the test subjects aside from those seen in the wild-type or vice versa.
Analysis
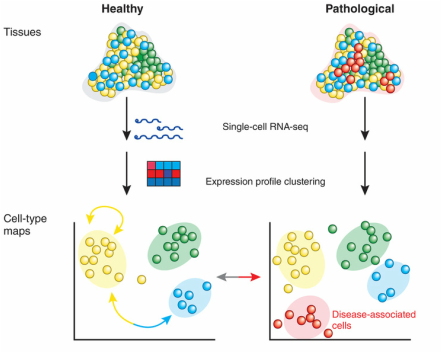
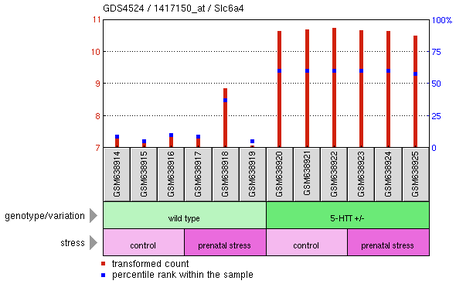
Gene Expression Omnibus (GEO) was used to find primary literature about the transcriptome of cells expressing SLC6A4. Most of the experiments on SLC6A4 were not related to anxiety or mood disorders of any kind, but one study by Hove, et al. looked at the effects of environmental stress on geneotypically heterozygous 5-Htt knockout mice, which would be analogous to SLC6A4 knockouts in humans. The methods, in brief, involved using 5-Htt +/- mice (only one chromosomal gene knocked out) to breed, and then exposing test females to environmental stresses throughout the pregnancy while leaving the control females to gestate as normal. Offspring where genotyped to see if they were 5-Htt +/+, +/-, or -/-. After two months, offspring were tested on a number of behavioral responses such as object recognition and forced swim tasks. The data from these tasks indicated that female 5-Htt +/- mice were most strongly affected, so they were studied further. Based on examination of their hippocampi, the graph below was able to be constructed.
Interpretation of the results shows that the degree to which prenatal stress affects a mouse is correlated to its genotype. Wild-type mice were not strongly effected by prenatal stress overall. However, the heterozygous 5-Htt +/- mice were strongly effected by prenatal stress and showed a greater amount of sensitivity to gene-environment interactions than the wild-type mice as well as stronger depression-like symptoms. The researchers went one step further and determined how certain genes were differentially expressed in each of the mouse combinations. GEO DataSets was used to compare the expression of those genes in each of the offspring to determine the mechanisms by which genotype and environmental stressors affect the transcriptome.
In this experiment, the 5-Htt and SLC6A4 genes are synonymous. Expression was found to be very high in the 5-Htt +/- mice, regardless of the environmental stressors applied to them, meaning there were lots of SLC6A4 proteins present and a high degree of serotonin reuptake occurring. These mice also showed higher rates of depression and anxious behaviors, which supports other primary literature that states low levels of serotonin lead to depression and anxiety disorders. Expression in all wild-type mice was low, suggesting more serotonin was present in the synapses of these mice. This would lead to elevated moods in these mice compared to the test mice and therefore lower rates of depression and anxiety disorders.
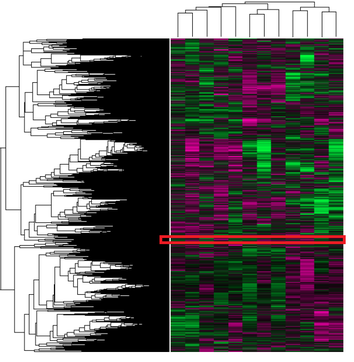
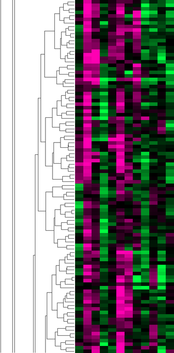
The 380 genes in the close-up of the microarray above were entered into GO for enrichment analysis to see if there were any related biological processes, cellular components, or molecular functions commonly found between them. The most common molecular function terms found were very general, indicating there was not a high degree of similarity in the characteristics of the genes. The top three terms were binding (GO:0005488), protein binding (GO:0005515), and basal transcription machinery binding (GO:0001098). These are terms that could be associated with almost any gene. Similarly broad terms were found for biological processes (regulation of metabolic process GO:0019222, biological regulation GO:0065007, cellular process GO:0009987, single-organism process GO:0044699) and cellular components (cell GO:0005623, intracellular part GO:0044424). The only conclusion that can be that there is not a strong enough relationship between the genes in this part of the transcriptome to represent a more detailed GO term vocabulary.
The 380 genes in the close-up of the microarray above were entered into GO for enrichment analysis to see if there were any related biological processes, cellular components, or molecular functions commonly found between them. The most common molecular function terms found were very general, indicating there was not a high degree of similarity in the characteristics of the genes. The top three terms were binding (GO:0005488), protein binding (GO:0005515), and basal transcription machinery binding (GO:0001098). These are terms that could be associated with almost any gene. Similarly broad terms were found for biological processes (regulation of metabolic process GO:0019222, biological regulation GO:0065007, cellular process GO:0009987, single-organism process GO:0044699) and cellular components (cell GO:0005623, intracellular part GO:0044424). The only conclusion that can be that there is not a strong enough relationship between the genes in this part of the transcriptome to represent a more detailed GO term vocabulary.
References
[1] Sandberg, R. (2013). Entering the era of single-cell transcriptomics in biology and medicine. Nature Methods, 11, 22-24. Retrieved March 24, 2015, from http://www.nature.com.ezproxy.library.wisc.edu/nmeth/journal/v11/n1/full/nmeth.2764.html
[2] Hove, D., Jakob, S., Schraut, K., Kenis, G., Schmitt, A., Kneitz, S., ... Burne, T. (2011). Differential Effects of Prenatal Stress in 5-Htt Deficient Mice: Towards Molecular Mechanisms of Gene × Environment Interactions. PLoS ONE, 6(8), E22715-E22715. Retrieved March 24, 2015, from http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3155516/
Gene Expression Omnibus: http://www.ncbi.nlm.nih.gov/geoprofiles
GEO DataSets: http://www.ncbi.nlm.nih.gov/sites/GDSbrowser?acc=GDS4524
GeneOntology: http://geneontology.org/
Pictures from top to bottom:
http://www.nature.com/nbt/journal/v30/n8/fig_tab/nbt.2325_F1.html (with modifications made in Paint)
http://www.nature.com.ezproxy.library.wisc.edu/nmeth/journal/v11/n1/full/nmeth.2764.html
http://www.ncbi.nlm.nih.gov/geo/tools/profileGraph.cgi?ID=GDS4524:1417150_at
http://www.ncbi.nlm.nih.gov/geo/gds/analyze/analyze.cgi?ID=GDS4524
http://www.ncbi.nlm.nih.gov/geo/gds/analyze/analyze.cgi?ID=GDS4524 (close-up)
[2] Hove, D., Jakob, S., Schraut, K., Kenis, G., Schmitt, A., Kneitz, S., ... Burne, T. (2011). Differential Effects of Prenatal Stress in 5-Htt Deficient Mice: Towards Molecular Mechanisms of Gene × Environment Interactions. PLoS ONE, 6(8), E22715-E22715. Retrieved March 24, 2015, from http://www.ncbi.nlm.nih.gov/pmc/articles/PMC3155516/
Gene Expression Omnibus: http://www.ncbi.nlm.nih.gov/geoprofiles
GEO DataSets: http://www.ncbi.nlm.nih.gov/sites/GDSbrowser?acc=GDS4524
GeneOntology: http://geneontology.org/
Pictures from top to bottom:
http://www.nature.com/nbt/journal/v30/n8/fig_tab/nbt.2325_F1.html (with modifications made in Paint)
http://www.nature.com.ezproxy.library.wisc.edu/nmeth/journal/v11/n1/full/nmeth.2764.html
http://www.ncbi.nlm.nih.gov/geo/tools/profileGraph.cgi?ID=GDS4524:1417150_at
http://www.ncbi.nlm.nih.gov/geo/gds/analyze/analyze.cgi?ID=GDS4524
http://www.ncbi.nlm.nih.gov/geo/gds/analyze/analyze.cgi?ID=GDS4524 (close-up)
|
GO Terms:
binding (GO:0005488) protein binding (GO:0005155) basal transcription machinery binding (GO:0001098) regulation of metabolic process (GO:0019222) |
biological regulation (GO:0065007)
cellular process (GO:0009987) single-organism process (GO:0044699) cell (GO:0005623) intracellular part (GO:0044424) |